Unidades Centrales de Apoyo a la Investigación Biomédica
Unidad de Espectrometría de Masas e Imagen Molecular de IMIBIC (IMSMI)

La Unidad de Espectrometría de Masas e Imagen Molecular de IMIBIC (IMSMI) ofrece soporte altamente especializado en las últimas técnicas de espectrometría de masas disponibles a aquellos investigadores y usuarios que así lo requieran. Esta Unidad, está especializada en ofrecer servicios en tres de las grandes -ómicas como son Proteómica, Lipidómica y Metabolómica. Además, está particularmente especializada en Imagen Molecular (MS-Imaging).
IMSMI ofrece sus servicios tanto a los grupos de investigación de IMIBIC, de la Universidad de Córdoba, del Hospital Universitario Reina Sofía, como de otras universidades, hospitales y compañías del sector privado.
La unidad posee cuatro líneas de trabajo principales:
- i) Línea de Proteómica Clínica, compuesta por un cromatógrafo líquido de alto rendimiento acoplado a espectrometría de masas (LC-MS/MS). Proporciona a los investigadores y usuarios el acceso a análisis de Proteómica Clínica cuantitativa en líquido y sin marcaje (DDA, DIA y PRM). Los estudios que pueden desarrollarse en esta línea van desde pequeños experimentos de pilotaje o pequeñas poblaciones a grandes experimentos con cientos de muestras y que necesiten de una velocidad y calidad superior.
- ii) Línea de Lipidómica, compuesta por un cromatógrafo líquido acoplado a espectrometría de masas (LC-MS/MS). Presta servicio de identificación y cuantificación de especies lipídicas en muestras de manera dirigida (targeted) o no dirigida (untargeted) a alta resolución.
- iii) Línea de Metabolómica, formada por un cromatógrafo líquido acoplado a espectrometría de masas (LC-MS/MS). Ofrece servicio de identificación y cuantificación de metabolitos en muestras de manera dirigida (targeted) o no dirigida (untargeted) a alta resolución.
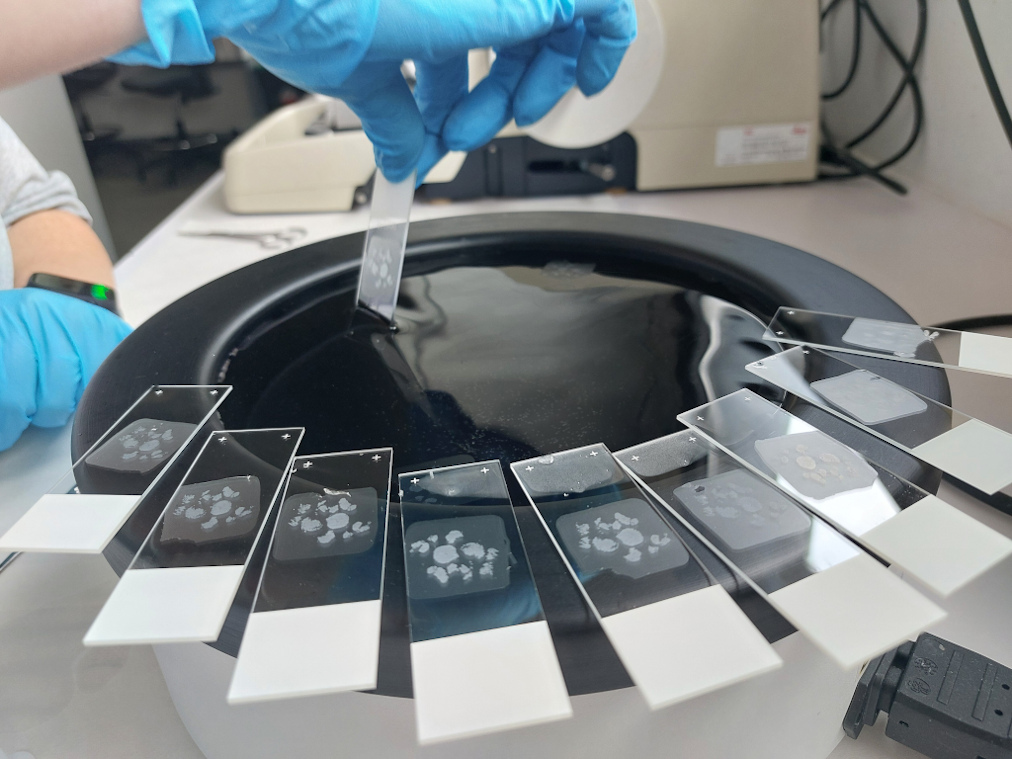
- iv) Línea de Imagen Molecular (MS-Imaging), que provee a los investigadores y usuarios información espacial sobre metabolitos, lípidos, proteínas/péptidos directamente de tejidos y biopsias.
Además, IMSMI proporciona soporte a sus usuarios de manera individualizada y personalizada incluyendo en el mismo el diseño experimental, preparación de muestra, análisis mediante espectrometría de masas y análisis de datos tanto básico como avanzado, incluyendo los últimos avances disponibles en su área de conocimiento.
La unidad de Espectrometría de Masas e imagen Molecular dispone de un Sistema de Gestión de Calidad conforme a la Norma de Calidad ISO 9001:2015.
Personal

Eduardo Chicano Gálvez, PhD
Responsable de la Unidad. Especialista Técnico Senior
- eduardo.chicano@imibic.org
- +34 957 213 757 (Corporativo 3757)

Ángela Peralbo Molina, PhD
Especialista Técnico Senior
- angela.peralbo@imibic.org
- +34 957 213 754 (Corporativo 3754)

Cristina Mª López Vázquez
Especialista Técnico Junior
- cristina.lopez@imibic.org
- +34 957 213 757 (Corporativo 3757)

Ana Salinas Gavilán
Técnica de laboratorio
- ana.salinas@imibic.org
- +34 957 213 757 (Corporativo 3757)
Equipamiento e instalaciones
- NanoELUTE2 (Bruker) (Last update: 2024)
- Beatbox (Preomics) (Last update: 2024)
- ThermoScientific Savant SpeedVac (Last update: 2025)
- Espectrómetro de masas QTOF, MALDI-TOF, TIMSTOF-Flex (Bruker) (Última actualización: 2022)
- Espectrómetro de masas Q-TOF, TIMSTOF-Pro (Bruker) (Última actualización: 2021)
- EVOSEP-ONE (Evosep) (Last update: 2021)
- ELUTE UHPLC (Bruker) (Last update: 2021)
- Sprayer: HTX M5 (HTX Technologies) (Last update: 2021)
- Principales programas (y lenguajes de programación) utilizados para el análisis de datos: TransProteomicsPipeline, Fragpipe for protein identification, Skyline, DIANN, MS-Fragger for protein quantification and spectral library generation, Cardinal, SCILS 2026b for Maldi Imaging MS analysis. R, Python. MetaboScape and TASQ for metabolomics/lipidomics workflow (Última actualización: 2026)
- BPS: Bruker Proteoscape. Search Engine and QC in Real-time (Bruker) (Last update: 2026)
- Equipamiento básico para preparación de muestras (Last update: 2021)
- Spectronaut 20 (Biognosys) (Last update: 2026)
Servicios
- Identificación y caracterización de proteínas mediante LC-MS/MS: DDA-PASEF
- Proteómica Cuantitativa:
- DIA-PASEF
- DIA-SWATH
- PRM-PASEF
- SRM (Selected Reaction Monitoring, QqQ)
- Clinical Proteomics analysis using Evosep-One
- Lipidómica y Metabolómica: Identificación, caracterización y/o cuantificación de pequeña molécula en un amplio rango de muestras
- Imagen Molecular (MSImaging (Proteómica/Lipidómica))
- Supervisión de proyectos. Colaboraciones.
- Formación en software utilizado para el análisis de datos
Publicaciones más relevantes
2026
2025
- Identification of diagnostic biomarkers for relapsing-remitting multiple sclerosis in plasma by mass spectrometry-based proteomics. Víctor M. García-Silva, Eduardo Chicano-Galvez, María Isabel García-Sánchez, Ángela Peralbo-Molina, María Hernández-Valladares. Journal of Neuropathology & Experimental Neurology, 2025, 1–9. https://doi.org/10.1093/jnen/nlaf145
- Proteomic Validation of MEG-01-Derived Extracellular Vesicles as Representative Models for Megakaryocyte- and Platelet-Derived Extracellular Vesicles. Jose Manuel Sanchez-Manas, Sonia Perales, Gonzalo Martinez-Navajas, Jorge Ceron-Hernandez, Cristina M. Lopez, Angela Peralbo-Molina, Juan R. Delgado, Joaquina Martinez-Galan, Veronica Ramos-Mejia, Eduardo Chicano-Galvez, Maria Hernandez-Valladares, Francisco M. Ortuno, Carolina Torres and Pedro J. Real. Biomolecules 2025, 15(12), 1968. https://doi.org/10.3390/biom15121698
- A Compendium of Bona Fide Reference Markers for Genuine Plant Extracellular Vesicles and Their Degree of Phylogenetic Conservation. Miriam M. Rodríguez de Lope, Ibone R. Sánchez-Pajares, Estela Herranz, Cristina M. López-Vázquez, Ainara González-Moro, Alan Rivera-Tenorio, Carlos González-Sanz, Soledad Sacristán, Eduardo Chicano-Gálvez, Fernando de la Cuesta. Journal of Extracellular Vesicles, 2025; 14:e70147. https://doi.org/10.1002/jev2.70147
- Defective Olfactomedin-2 connects adipocyte dysfunction to obesity. Aina Lluch, Jèssica Latorre, Isabel Espadas, Núria Oliveras-Cañellas, José M. Moreno-Navarrete, Estefanía Caballano-Infantes, Gitalee Sarker, Nicolás F. Malvido, Pablo Garrido-Gil, José L. Labandeira-García, Naoki Nakaya, Silvia Mora, Eduardo Chicano, Jaime López-Alcalá, María M. Malagón, Alejandro Martín-Montalvo, Birong Zhang, You Zhou, Ana I. Domingos, Miguel López, Johanna Pörschke, María Gómez-Serrano, Witold Szymanski, Johannes Graumann, Stanislav I. Tomarev, Ismael González-García, José M. Fernández-Real, Francisco J. Ortega. Nature Communications | (2025) 16:7154. https://doi.org/10.1038/s41467-025-62430-5
- Spatially resolved lipid disruption induced in Daphnia magna by environmentally relevant exposure to obesogens tributyltin and pyriproxyfen. Albert Menéndez-Pedriza, Cristina-María López, Eduardo Chicano-Gálvez, Joaquim Jaumot, Carlos Barata, Laia Navarro-Martín. Aquatic Toxicology 287 (2025) 107521
- Comprehensive MSI-based protocols for the spatial lipidomics characterization of microscale organisms. Albert Menéndez-Pedriza, Christoph Bookmeyer, Michiel Vandenbosch, Sutirtha Chattopadhyay, María García-Altares, Eduardo Chicano-Gálvez, Laia Navarro-Martín, Joaquim Jaumot. Microchemical Journal 2025. https://doi.org/10.1016/j.microc.2025.114936
- Afamin and Apolipoprotein F Associated With Liver Steatosis From People Living With HIV: A Discovery Study. Mario Frías, Eduardo Chicano-Gálvez, Antonio Rivero-Juárez, Ana Gordon, Diana Corona-Mata, José María Moyano, Ángela Peralbo-Molina, Ángela Camacho, Ignacio Pérez-Valero, María del Mar Malagón, Antonio Rivero. Alimentary Pharmacology and Therapeutics, 2025. https://doi.org/10.1111/apt.70119
- A MALDI-MSI-based approach to characterize the spatial distribution of cylindrospermopsin and lipid alterations in rat intestinal tissue. Antonio Casas-Rodríguez, Cristina María López-Vázquez, Remedios Guzmán-Guillén, Nahúm Ayala, Ana María Cameán, Angeles Jos, Eduardo Chicano-Gálvez. Chemico-Biological Interactions 412 (2025) 111479. https://doi.org/10.1016/j.cbi.2025.111479
- Quantitative proteomic analysis unveils a critical role of VARS1 in hepatocellular carcinoma aggressiveness through the modulation of MAGI1 expression. Natalia Hermán-Sánchez, Mercedes del Rio-Moreno, Rubén Ciria, Marina E. Sánchez-Frias, Maite G. Fernández-Barrena, Iker Uriarte, Eduardo Chicano-Galvez, Ignacio Ortea, Ángela Peralbo-Molina, Javier Briceño, Matías A. Avila, Manuel Rodríguez-Perálvarez, Raúl M. Luque, Juan L. López-Cánovas & Manuel D. Gahete. Molecular Cancer 24, 15 (2025). https://doi.org/10.1186/s12943-024-02206-5
2024
- Porin expression in clinical isolates of Klebsiella pneumoniae: a comparison of SDS-PAGE and MALDI-TOF/MS and limitations of whole genome sequencing analysis. Cristina Elías-López, Montserrat Muñoz-Rosa, Julia Guzmán-Puche, Elena Pérez-Nadales, Eduardo Chicano-Galvez, Luis Martínez-Martínez. Annals of Clinical Microbiology and Antimicrobials (2024).
- Longitudinal Assessment of Nasopharyngeal Biomarkers Post-COVID-19: Unveiling Persistent Markers and Severity Correlations. Francisco Javier Redondo-Calvo, Yoana Rabanal-Ruiz, Gema Verdugo-Moreno, Natalia Bejarano-Ramírez, Raquel Bodoque-Villar, Mario Durán-Prado, Soledad Illescas, Eduardo Chicano-Galvez, Francisco Javier Gómez-Romero, José Martinez-Alarcón, Javier Arias-Pardilla, Pilar Lopez-Juarez, Juan Fernando Padin, Juan Ramón Peinado, Leticia Serrano-Oviedo. Journal of Proteome Research (2024).
- Superior metabolic improvement of polycystic ovary syndrome traits after GLP1-based multi-agonist therapy. Miguel A. Sánchez-Garrido, Víctor Serrano-López, Francisco Ruiz-Pino, María Jesús Vázquez, Andrea Rodríguez-Martín, Encarnación Torres, Inmaculada Velasco, Ana Belén Rodríguez, Eduardo Chicano-Gálvez, Marina Mora-Ortiz, Claes Ohlsson, Matti Poutanen, Leonor Pinilla, Francisco Gaytán, Jonathan D. Douros, Bin Yang, Timo D. Müller, Richard D. DiMarchi, Matthias H. Tschöp, Brian Finan, Manuel Tena-Sempere . Nature Communications 15, 8498 (2024)
- Pollen-Food Allergy Syndrome: From Food Avoidance to Deciphering the Potential Cross-Reactivity between Pru p 3 and Ole e 7. Paula Álvarez,Rocío Aguado, Juan Molina, Antonio Trujillo-Aguilera, Mayte Villalba, Araceli Díaz-Perales, Carmen Oeo-Santos, Eduardo Chicano, Nadine Blanco, Ana Navas, Berta Ruiz-León, Aurora Jurado. Nutrients 2024, 16(17), 2869
- Dysregulation of Lipid Metabolism Serves as A Link Between Alzheimer’s and Cardiovascular Disease, As Witnessed in A Cross-Sectional Study. Laura Mourino-Alvarez, Cristina Juarez-Alia, Tamara Sastre-Oliva, Inés Perales-Sánchez, German Hernandez-Fernandez, Eduardo Chicano-Galvez, Ángela Peralbo-Molina, Felipe Madruga, Emilio Blanco-Lopez, Teresa Tejerina, María G. Barderas. Aging and disease. Volume 16, Number 3, June 2024.
- Revisiting tuberculous granuloma by MALDI-Imaging Proteomic analysis of granulomas from cattle and pigs naturally infected with Mycobacterium tuberculosis complex by MALDI-imaging. Fernanda I. Larenas-Muñoz, José María Sánchez-Carvajal, Inés Ruedas-Torres, Carmen Álvarez-Delgado, Karola Fristiková, Francisco J. Pallarés, Librado Carrasco, Eduardo Chicano-Gálvez, Irene Magdalena Rodríguez-Gómez, Jaime Gómez-Laguna. Frontiers in Immunology. Volume 15 - 2024 | doi: 10.3389/fimmu.2024.1369278
- Mass spectrometry imaging in environmental monitoring: From a scarce existing past to a promising future. Ana María Herruzo-Ruiz, Angela Peralbo-Molina, Cristina-María López, Carmen Michán, José Alhama, Eduardo Chicano-Gálvez. Trends in Environmental Analytical Chemistry. Volume 42, June 2024. https://doi.org/10.1016/j.teac.2024.e00228
2023
- Development of a Prediction Modelo for Short-Term Remission of Patients with Crohn's Disease Treated with Anti-TNF Drugs. Rosario Medina-Medina et al. Int. J. Mol. Sci. 2023, 24(10), 8695; https://doi.org/10.3390/ijms24108695
- Development of a Prediction Model for Short-Term Remission of Patiens with Crohn's Disease Treated with Anti-TNF Drugs. Rosario Medina-Medina, Eva Iglesias-Flores, Jose M. Benítez, Sandra Marín-Pedrosa, Isabel Salguerio-Rodríguez, Clara I. Linares, Sandra González-Rubio, Pilar Soto-Escribano, Beatriz Gros, Manuel L. Rodríguez-Perálvarez, José L. Cabriada, María Chaparro, Javier P. Gisbert, Eduardo Chicano-Gálvez, Ignacio Ortea, Gustavo Ferrín, Valle García-Sánchez and Patricia Aguilar-Melero. Int. J. Mol. Sci. 2023, 24(20), 8695. May 12, 2023. https://doi.org/10.3390/ijms24108695
- The thrombus proteome in stroke reveals a key role of the innate immune system and new insights associated with its aetiology, severity and prognosis. Chary Lopez-Pedrera, Rafael Oteros, Alejandro Ibáñez-Costa, María Luque-Tévar, Laura Muñoz-Barrera, Nuria Barbarroja, Eduardo Chicano-Gálvez, Juan Marta-Enguita, Josune-Orbe, Francisco Velasco, Carlos Perez-Sánchez. Journal of thrombosis and haemostasis. April 24, 2023 DOI:https://doi.org/10.1016/j.jtha.2023.04.015
- Spatial and Temporal Protein Modules Signatures Associated with Alzheimer Disease in 3xTg-AD Mice Are Restored by Early Ubiquinol Supplementation. Emilio Llanos-González , Francisco J. Sancho-Bielsa, Javier Frontiñán-Rubio, Yoana Rabanal-Ruíz, Sonia García-Carpintero, Eduardo Chicano, Isabel Úbeda- Banon, Alicia Flores-Cuadrado, Lydia Giménez-Llort, Francisco Javier Alcaín, Juan Ramón Peinado and Mario Durán-Prado. Antioxidants 2023, 12(3), 747; https://doi.org/10.3390/antiox12030747
- Data Processing and Analysis in Mass Spectrometry-Based Metabolomics. Ángela Peralbo-Molina, Pol Solà-Santos, Alexandre Perera-Lluna, Eduardo Chicano-Gálvez. Methods Mol Biol. 2023;2571:207-239. doi: 10.1007/978-1-0716-2699-3_20
2022
- Data Processing and Analysis in Mass Spectrometry-Based Metabolomics. Ángela Peralbo-Molina, Pol Solà-Santos, Alexandre Perera-Lluna & Eduardo Chicano-Gálvez. Mass Spectrometry for Metabolomics. Methods in Molecular Biology, vol 2571. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2699-3_20
- The NtrYX Two-Component System of Paracoccus denitrificans Is Required for the Maintenance of Cellular Iron Homeostasis and for a Complete Denitrification under Iron-Limited Conditions. Alfonso Olaya-Abril, Víctor M Luque-Almagro, Jesús Hidalgo-Carrillo, Eduardo Chicano-Gálvez, Francisco J Urbano, Conrado Moreno-Vivián, David J Richardson, María Dolores Roldán. Int J Mol Sci. 2022 Aug 15;23(16):9172. doi: 10.3390/ijms23169172.
2021
2020
2019
- González-Fernández MJ, Fabrikov D, Ramos-Bueno RP, Guil-Guerrero JL, Ortea I.SWATH Differential Abundance Proteomics and Cellular Assays Show in Vitro Anticancer Activity of Arachidonic Acid-and Docosahexaenoic Acid-Based Monoacylglycerols in HT-29 Colorectal Cancer Cells. Nutrients 2019, 11, 2984
- López-Sánchez LM, Jiménez Izquierdo R, Peñarando J, Mena R, Guil-Luna S, Toledano M, Conde F, Villar C, Díaz C, Ortea I, De la Haba-Rodríguez JR, Aranda E, Rodríguez-Ariza A.SWATH-based proteomics reveals processes associated with immune evasion and metastasis in poor prognosis colorectal tumours. J Cell Mol Med.2019, 23, 8219-8232
- Michán C, Chicano-Gálvez E, Fuentes-Almagro CA, Alhama J. Redox and global interconnected proteome changes in mice exposed to complex environmental hazards surrounding Doñana National Park. Environ Pollut 2019, 252, 427-439
- Böhme K, Calo-Mata P, Barros-Velázquez J, Ortea I. Recent applications of omics-based technologies to main topics in food authentication. TrAC - Trends in Analytical Chemistry 2019, 110, 221-232.
- Böhme, K.; Calo-Mata, P.; Barros-Velázquez, J.; Ortea, I. Review of Recent DNA-Based Methods for Main Food-Authentication Topics. Journal of Agricultural and Food Chemistry 2019, 67, 3854-3864.
- Garrido-Rodríguez M, Ortea I, Calzado MA, Muñoz E, García V. SWATH proteomic profiling of prostate cancer cells identifies NUSAP1 as a potential molecular target for Galiellalactone. J Proteomics. 2019, 20;193:217-229.
2018
- Ortea I,
Ruiz-Sánchez I, Cañete R, Caballero-Villarraso J, Cañete MD. Identification of candidate serum
biomarkers of childhood-onset growth hormone deficiency using SWATH-MS and feature selection. J
Proteomics 2018, 20;175:105-113.
- Fernández A, Bayón GF, Sierra MI, Ortea I, Bonilla F, Fraga M. Loss of 5hmC identifies a new type of aberrant DNA hypermethylation in glioma. Hum Mol Genet. 2018 Sep 1;27(17):3046-3059.
- Ortea I, González-Fernández MJ, Ramos-Bueno RP, Guil-Guerrero JL. Proteomics Study Reveals That Docosahexaenoic and Arachidonic Acids Exert Different in Vitro Anticancer Activities in Colorectal Cancer Cells. J Agric Food Chem. 2018 Jun 20;66(24):6003-6012
2017
2016
- Prado M, Ortea I, Vial S, Rivas J, Calo-Mata P, Barros-Velázquez J. Advanced DNA- and protein-based methods for the detection and investigation of food allergens. Crit Rev Food Sci Nutr. 2016 Nov 17;56(15):2511-2542.
- Ortea I, Rodriguez-Ariza A, Chicano E, Arenas MS, Jurado B. Discovery of potential protein biomarkers of lung adenocarcinoma in bronchoalveolar lavage fluid by SWATH MS data-independent acquisition and targeted data extraction. J Proteomics. 2016 Apr 14;138:106-14.
- Ghedira J, Chicano-Gálvez E, Fernández-Cisnal R, Jebali J, Banni M, Chouba L, Boussetta H, López-Barea J, Alhama J. Using environmental proteomics to assess pollutant response of Carcinus maenas along the Tunisian coast. Sci Total Environ. 2016 Jan 15;541:109-118.
- Ortea I, Roschitzki B et al. Independent Candidate Serum Protein Biomarkers of Response to Adalimumab and to Infliximab in Rheumatoid Arthritis: An Exploratory Study. PLoS One. 2016 Apr 6;11(4):e0153140.
- Ortea I, O'Connor G, Maquet A. Review on Proteomics for food authentication. J Proteomics. 2016 Sep 16;147:212-22.
2015
- Chicano-Gálvez E, Asensio E, Cañavate JP, Alhama J, López-Barea J. Proteomic analysis through larval development of Solea senegalensis flatfish. Proteomics. 2015 Dec;15(23-24):4105-19.
- Abril N, Chicano-Gálvez E, Michán C, Pueyo C, López-Barea J. iTRAQ analysis of hepatic proteins in free-living Mus spretus mice to assess the contamination status of areas surrounding Doñana National Park (SW Spain). Sci Total Environ. 2015 Aug 1;523:16-27.
- de Andrés MC, Pérez-Pampín E, Calaza M, Santaclara FJ, Ortea I, Gómez-Reino JJ, González A. Assessment of global DNA methylation in peripheral blood cell subpopulations of early rheumatoid arthritis before and after methotrexate. Arthritis Res Ther. 2015 Aug 29;17:233